In the headlines
Become a curator of the site
This web site is the site of anyone interested in contributing to the promotion of Brucella genotyping. If you want to become a curator, and contribute to this new "MLVApedia" wiki just contact any of the current curators and join. By being a curator, you can also use this site to promote the work of your own laboratory in this field.
News
2016, May 16th, MicrobesGenotyping site released for beta-testing
The MicrobesGenotyping website software has been fully rewritten based on our previous experience with http://mlva.u-psud.fr. The Brucella database has been updated and is included in the new version. The database now contains data for more than 4000 strains.
Brucella2012 released
Brucella2012 has been released. It contains published MLVA data from more than 1400 isolates. The data can be accessed via the version 4 MLVA site. Important points: 1-the coding convention for Bruce19 is changing. Due to an internal 15 bp microdeletion in some rare alleles, the allele size distribution looks as if the repeat unit was 3 bp long. In order to cope with this, the coding convention has been changed as follows : initial convention : bruce19-BRU324_6bp_163bp_18u new convention (...)
Brucella2010 released
A new compilation of MLVA data has been released A new compilation of MLVA data has been released, under database name Brucella2010. It contains MLVA16 data from 707 isolates. Two preselected panels of loci are proposed in addition to the full 16 loci. MLVA8 alias panel1 corresponds to the 8 loci Bruce06, Bruce08, Bruce11, Bruce12, Bruce42, Bruce43, Bruce45, Bruce55. MLVA11 is the 11 loci including panel 1 plus 2A, i.e. Bruce18, Bruce19, Bruce21. The previous releases, Brucella2007 and (...)
Brucella2009 just released
The Brucella2009 MLVA-16 database with data from more than 500 isolates covering all known Brucella species and biovars was just released. This is a major update to the previous version Brucella2007. Brucella2009 compiles published data using the MLVA-16 assay. In particular, the recently published new Brucella species (microti, ceti, pinnipedialis) are covered in detail. We recommand the reading of the associated report by Maquart et al., 2009. The data in Brucella2009 is accessible as (...)

Diversity of clinical Brucella melitensis isolates in Lebanon
Kattar et al. report the investigation of the genetic diversity of 42 clinical Brucella melitensis isolates from Lebanon. Diversity was evaluated by MLVA analysis, and the genotypes observed are typical "East Mediterranean" profiles.
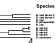
Brucella microti isolated from red fox in Austria
Scholz et al. report the isolation of Brucella microti from Austrian red foxes. The proposed hypothesis is that foxes could have become infected by ingestion of infected common voles.

Brucella microti isolated from soil
Scholz et al. report the isolation of Brucella microti in soil. Brucella microti is a recently described species within the Brucella genus. Its behaviour is strinkingly different in terms of growth characteristics (it is a fast growing microorganism) compared to the rest of the genus. The finding of this species in soil is in keeping with this observation, and provides a first indication of where Brucella as a highly host adapted pathogen may come (...)
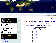
The Brucella2007 MLVA database just released
The latest version of the Brucella database has just been released. It compiles data published in Le Flèche et al., 2006, Al Dahouk et al., 2007, Garcia-Yoldi et al., 2007a and 2007b, Scholz et al., 2008, Kattar et al., 2008. The data is derived from 529 isolates, and using MLVA16 384 genotypes are resolved. More detailed support information can be found on the MLVA wiki and the database can be accessed at (...)

MLVA ring trial announcement : closing time
The first MLVA ring trial will be closed end of february 2008. All participants need to have send their results to Falk Melzer by then. As a reminder, the first ring trial is focused on panel 1. The results can be submitted as a table with repeat copy number for each sample. Help for allele assignment can be obtained from http://mlva.u-psud.fr/Brucella/ and a panel 1 genotype number can also be assigned using the public ressource at http://minisatellites.u-psud.fr/MLVAnet/ alias (...)

New species Brucella microti validly published
Brucella microti, a common vole Brucella species is published in the February 2008 issue of the IJSEM journal ; the name is placed on the ICSP List of Prokaryotic Names with Standing in Nomenclature and genus Brucella is now harbouring nine described species.

Two new species within the Brucella genus
The International Journal of Systematic and Evolutionary Microbiology published in the 2007 november issue (57, 2688–2693) Brucella ceti sp. nov. and Brucella pinnipedialis sp. nov. for Brucella strains with cetaceans and seals as their preferred hosts (G. Foster et al.). The names of Brucella ceti (cetaceans as preferred hosts) and Brucella pinnipedialis (seals as preferred hosts), attached to these two new described Brucella species (with type strains deposited at the NCTC and at the (...)

MLVA ring trial 2007, current list of participants
Twenty institutions from fifteen countries have registered to participate to the first Brucella MLVA ring trial. The distribution of samples has already begun (December, 4th). The samples should arrive within the next days. Kindly acknowledge receipt.
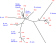
MLST investigation within the Brucella genus
Whatmore et al. have investigated the genetic diversity and phylogeny of the Brucella genus by multiple loci sequence typing. More than 4 kb of sequence were examined from 160 isolates. 27 sequence types can be distinguished. The clustering achieved confirms the close vicinity of B. canis with B. suis biovar 3 and 4, and the marked difference with B. suis biovar 5. The marine strains are tightly clustered by this approach. Although much less discriminatory than MLVA, MLST data provides a (...)
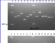

MLVA typing of Brucella isolates from humans in Sicily, Italy
MLVA-16 (i.e. Le Flèche 2006 + Bruce19, as described by Al Dahouk 2007) was applied to 20 human isolates from Sicily, Italy, by Marianelli et al.

MLVA analysis of Brucella isolates from human patients
Al Dahouk et al. published an MLVA investigation of Brucella isolates from human patients. One hundred and twenty-eight isolates were recovered from patients from France and Germany. Most cases were imported events, resulting from travels in mediterranean countries. MLVA patterns reflect to some extend the geographic origin of the isolates, when it can be deduced from the epidemiological investigation. The authors propose three main clusters, “East-Mediterranean”, “West-Mediterranean”, and (...)

Stability of MLVA loci in the Brucella melitensis Rev1 vaccine strain
Garcia-Yoldi et al. have used MLVA analysis to unambiguously identify Rev1 vaccine strain from field isolates. They used the Le Flèche 2006 panel of 15 loci. In some instances, variant alleles were observed at one of the most variable loci. This work is important to help define a strain in terms of MLVA typing.

MLVA for Brucella by Whatmore et al., 2006
Whatmore et al. published an MLVA assay comprising 21 loci. In this report, the length of fluorescently labelled PCR products is estimated by a capillary electrophoresis equipment. Nine loci are shared with the "Le Flèche" collection of markers.
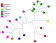
MLVA for Brucella by Le Flèche et al., 2006
Le Flèche et al published an MLVA assay for Brucella. It comprises 15 loci, and is the first MLVA assay for Brucella which is able to deduce the Brucella species from the MLVA genotype.
To be noted
2018, Brucella version 4_3 database update
Together with a significant upgrade of the MicrobesGenotyping web interface (version 1.4.0) the (...)
2016, May 19th, major Brucella database update
The Brucella database has been updated, it now contains genotyping data from over 4000 strains (...)
Second MLVA ring trial : join now !
Dear colleagues, after the great success of our first international MLVA ring trial we are now (...)
Brucellosis 2008
The Brucellosis 2008 International Conference (Including the 61st Brucellosis Research (...)
First MLVA ring trial : join now !
Participate to the first MLVA ring trial.
